Tag-accessed Memory for Genetic Programming
This repository is associated with our 2019 GECCO extended abstract submission, Tag-accessed Memory for Genetic Programming.
Feel free to contact us with questions or file an issue on this repository if something isn’t clear!
Navigation
Project Overview
We present an early exploration of tag-accessed memory for genetic programming.
Tag-accessed Memory
Tags are evolvable labels that give genetic programs a flexible mechanism for specification. Tag-based naming schemes have been demonstrated for labeling and referencing program modules (Spector, 2011; Lalejini and Ofria, 2018).
We continue to expand the use of tags in GP by incorporating tag-based referencing into the memory model of a simple linear GP representation. In this study, memory comprises 16 statically tagged memory registers, and instructions use tag-based referencing to refer to positions in memory. Programs in our simple representation are linear sequences of instructions, and each instruction has three tag-based arguments, which may modify the instruction’s behavior. Below, we provide a visual example, contrasting traditional direct-indexed memory access and tag-based memory access.
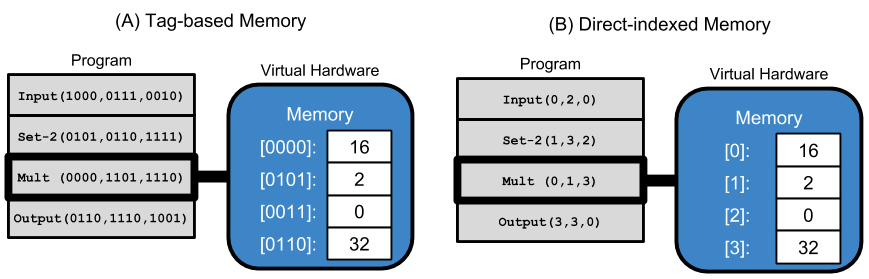
In the example above, both programs have identical behavior: requesting input, setting the second register to the terminal value ‘2’, multiplying the input by 2, and outputting the result.
Contribution Authors
Repository Guide
- analysis/
- Contains R scripts used for data analyses and generating graphs.
- data/
- Contains raw data for preliminary and published experiments as well as the training and testing examples used for the programming synthesis benchmark problems (taken from Tom Helmuth’s example repository).
- docs/
- Contains miscellaneous documentation associated with this work.
- experiment/
- Contains the source code (C++) for our simple linear GP representation and for running the experiments discussed in our contribution.
- hpcc/
- Contains scripts used to submit experiment jobs to MSU’s HPCC.
- media/
- Contains media (images) associated with this work.
- scripts/
- Contains utility scripts used for managing experiments on the HPCC and for aggregating and manipulating experiment data.
Supplemental Material
- Experiment configuration and GP system details: ./docs/gp-system.md
- Data analysis: Our analyses were done in R (R Core Team, 2016).
- Find a webpage (generated with R markdown) here: http://lalejini.com/GECCO-2019-tag-accessed-memory/analysis/tag-mem-analysis.html
- Or, the Rmd file is here: ./analysis/tag-mem-analysis.Rmd
References
Lalejini, A., & Ofria, C. (2018). Evolving event-driven programs with SignalGP. In Proceedings of the Genetic and Evolutionary Computation Conference on - GECCO ’18 (pp. 1135–1142). New York, New York, USA: ACM Press. https://doi.org/10.1145/3205455.3205523
R Core Team (2016). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL https://www.R-project.org/.
Spector, L., Martin, B., Harrington, K., & Helmuth, T. (2011). Tag-based modules in genetic programming. In Proceedings of the 13th annual conference on Genetic and evolutionary computation - GECCO ’11 (p. 1419). New York, New York, USA: ACM Press. https://doi.org/10.1145/2001576.2001767